Chronogenetics Research Group
Research topic:
Eukaryotic circadian clocks are oscillating gene networks, where the primary rhythm is generated at the level of transcription and is then adjusted by several additional mechanisms in order to keep the app. 24 h period (Figure 1.). This primary oscillation rhythmically regulates numerous molecular and physiological processes in a way that they appear at the most appropriate time of the day. Evidently, the clock fulfils its biological role only if its phase is synchronized to the environmental day/night cycles, and runs with a period close to 24 h.
Working with the model plant of modern plant biology, Arabidopsis thaliana, we characterized the light sensing proteins (photoreceptors) that absorb environmental light signals and described the transcription factors that transduce photoreceptor-mediated signals to the level of the regulation of clock gene expression, thus establishing the synchrony between the clock and the real world.
After performing a successful mutant screen in Arabidopsis thaliana, we identified mutations in two proteins pointing to novel regulatory modes of the plant circadian clock. The first protein is an ubiquitin protease (UBP12) removing ubiquitin moieties from proteins, including clock proteins. The specific mutation that we identified suggested that this function of the protease is regulated by phosphorylation. This hypothesis will be verified by the creating and analysing artificial derivatives fixed in the non/phosphorylated states. Homologues of UBP12 are present in animal systems, where they regulate the pace of the clock as well. Thus we also aim at testing if phosphorylation at the identified specific amino acid has similar effects on the circadian function of UBP12-like proteases in plants and animals.
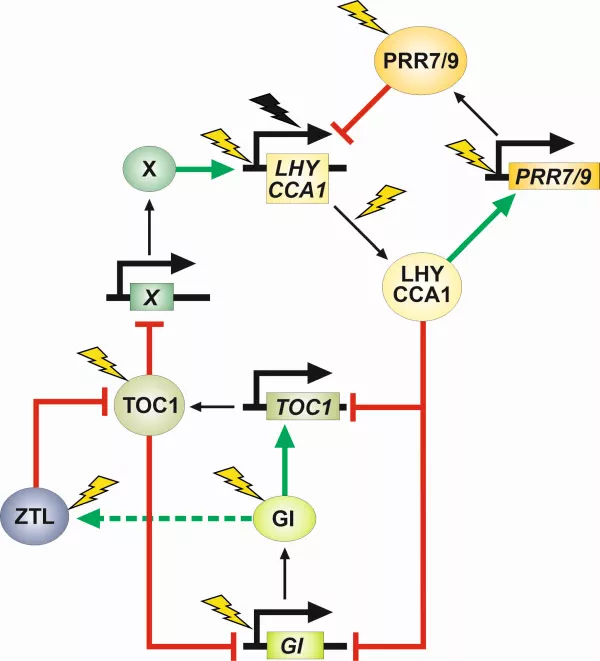
Figure 1. The structure of the plant circadian clock
Clock genes and clock proteins are indicated by rectangles and ellipses, respectively. Positive and negative regulatory effects are shown by green and red lines, respectively. Thunder symbols indicate light regulated steps (yellow: activation, black: inhibition). The plant clock is composed of three interlocked regulatory loop and their coordinated action is required to generate the primary 24 h oscillation. Although with altered period and amplitude, this robust regulatory system is capable of producing self-sustained oscillations even in the absence of one of the loops (e.g. because of the mutation of a particular clock gene)
The other protein has a well-known general interacting domain at the N-terminal part, whereas the mutation we identified marks a plant specific domain at the C-terminus with a yet unknown function. We aim to reveal this function by determining the set of clock and non-clock genes/proteins that are directly or indirectly regulated by this protein and by describing the mechanism of this regulation.
We continuously improve our research infrastructure and methodology covering numerous areas of modern molecular biology (analysis of gene expression, genetic mapping, gene isolation/modification, use of clock-regulated luminescent markers in transgenic organisms)(Figure 2.). It follows that skills and expertise acquired by using these procedures will be beneficial not only for plant chronobiologists, but also for researchers working on different fields or with different model organisms.
Circadian clocks regulate optimal growth of plants by orchestrating vital processes according to the day/night cycles. It is conceivable that the dramatic change in global climate demands re-organization of daily rhythms in such a fast manner, which cannot be matched by evolutionary processes. Under these circumstances, the knowledge of regulatory mechanisms that could be used to slightly change the phase of the clock within the day, but without affecting other fundamental processes in plants, will be most valuable.
Figure 2. Rhythmic expression of a clock-controlled promoter:luciferase gene construct in transgenic plants
The upper part of the figure shows a series of digital images taken of the same plants at different times. The time (h) and the day of taking a particular image are shown at the top and on the left side of the panel, respectively. The lower part of the figure shows a graphs where quantified luminescence values were plotted against the time. Light conditions are indicated by white (light) or black (dark) bars drawn over the time axis.
Publications
Merai Zsuzsanna, Xu Fei, Hajdu Anita, Kozma-Bognar Laszlo, Dolan Liam (2025). Phytochrome A is required for light-inhibited germination of Aethionema arabicum seed. NEW PHYTOLOGIST Read it
Hajdu Anita, Nyári Dóra Vivien, Ádám Éva, Kim Yeon Jeong, Somers David E., Silhavy Dániel, Nagy Ferenc, Kozma-Bognár László (2024). Forward genetic approach identifies a phylogenetically conserved serine residue critical for the catalytic activity of UBIQUITIN-SPECIFIC PROTEASE 12 in Arabidopsis. SCIENTIFIC REPORTS Read it
Hajdu Anita, Nyári Dóra, Terecskei Kata, Gyula Péter, Ádám Éva, Dobos Orsolya, Mérai Zsuzsanna, Kozma-Bognár László (2024). LIP1 Regulates the Plant Circadian Oscillator by Modulating the Function of the Clock Component GIGANTEA. CELLS Read it
Peter Csaba, Adam Eva, Klose Cornelia, Grezal Gabor, Hajdu Anita, Steinbach Gabor, Kozma-Bognar Laszlo, Silhavy Daniel, Nagy Ferenc, Viczian Andras (2024). Phytochrome C and Low Temperature Promote the Protein Accumulation and Red-Light Signaling of Phytochrome D. PLANT AND CELL PHYSIOLOGY Read it
Gombos Magdolna, Hapek Nóra, Kozma-Bognár László, Grezal Gábor, Zombori Zoltán, Kiss Edina, Györgyey János (2023). Limited water stress modulates expression of circadian clock genes in Brachypodium distachyon roots, SCIENTIFIC REPORTS Read it
Bernula Péter, Pettkó-Szandtner Aladár, Hajdu Anita, Kozma-Bognár László, Josse Eve-Marie, Ádám Éva, Nagy Ferenc, Viczián András (2021). SUMOylation of PHYTOCHROME INTERACTING FACTOR 3 promotes photomorphogenesis in Arabidopsis thaliana. NEW PHYTOLOGIST Read it
Kovacs Hajnalka, Aleksza David, Baba Abu Imran, Hajdu Anita, Kiraly Anna Maria, Zsigmond Laura, Toth Szilvia Z., Kozma-Bognar Laszlo, Szabados Laszlo (2019). Light Control of Salt-Induced Proline Accumulation Is Mediated by ELONGATED HYPOCOTYL 5 in Arabidopsis. FRONTIERS IN PLANT SCIENCE Read it
Gould PD, Domijian M, Greenwood M, Tokuda IT, Rees H, Kozma-Bognar L, Hall AJW, Locke JCW (2018). Coordination of robust single cell rhythms in the Arabidopsis circadian clock via spatial waves of gene expression. ELIFE Read it
Hajdu Anita, Dobos Orsolya, Domijan Mirela, Balint Balazs, Nagy Istvan, Nagy Ferenc, Kozma-Bognar Laszlo (2018). ELONGATED HYPOCOTYL 5 mediates blue light signalling to the Arabidopsis circadian clock. PLANT JOURNAL Read it